SBML Solver
When you study biology, sooner or later, you encounter pathway diagrams,
gene expression networks, Physiologicaly Based Pharmacokinetics (PBPK)
whole body diagrams, etc… Often, these can be mathematically represented
in the form of Ordinary Differential Equations (ODEs). There are many
ODE solvers available and you can write your own. However the solution
we like most is called SBML Solver. Before going any further let us
explain briefly what SBML itself is. SBML stands for Systems Biology Markup Language.
It was proposed around year 2000 by a few scientists from
Caltech (Mike Hucka, Herbert Sauro, Andrew Finney). According to
Wikipedia, SBML is a representation format, based on XML, for
communicating and storing computational models of biological processes.
In practice SBML focuses on reaction kinetics models but can also be
used to code these models that can be described in the form of ODEs such
as e.g. PBPK, population models etc…
Being a multi-cell modeling platform CC3D allows users to associate
multiple SBML model solvers with a single cell or create “free floating”
SBML model solvers. The CC3D Python syntax that deals with the SBML
models is referred to as “SBML Solver”. Internally SBML Solver relies on
C++ libRoadRunner developed by Herbert Sauro’s team. libRoadRunner in turn
is based on the C# code written by Frank Bergmann. CC3D uses
libRoadRunner as the engine to solve systems of ODEs. All SBML Solver
functionality is available via SteppableBasePy member functions.
Twedit++ provides nice shortcuts that help users write valid code while
tapping into SBML Solver functionality. See CC3DPython->SBML Solver menu
for options available.
Let us look at the example steppable that uses SBML Solver:
from cc3d.core.PySteppables import *
class SBMLSolverSteppable(SteppableBasePy):
def __init__(self, frequency=1):
SteppableBasePy.__init__(self, frequency)
def start(self):
# adding options that setup SBML solver integrator
# these are optional but useful when encountering integration instabilities
options = {'relative': 1e-10, 'absolute': 1e-12}
self.set_sbml_global_options(options)
model_file = 'Simulation/test_1.xml'
initial_conditions = {}
initial_conditions['S1'] = 0.00020
initial_conditions['S2'] = 0.000002
self.add_sbml_to_cell_ids(model_file=model_file, model_name='dp',
cell_ids=list(range(1, 11)), step_size=0.5, initial_conditions=initial_conditions)
self.add_free_floating_sbml(model_file=model_file, model_name='Medium_dp', step_size=0.5,
initial_conditions=initial_conditions)
self.add_free_floating_sbml(model_file=model_file, model_name='Medium_dp1', step_size=0.5,
initial_conditions=initial_conditions)
self.add_free_floating_sbml(model_file=model_file, model_name='Medium_dp2')
self.add_free_floating_sbml(model_file=model_file, model_name='Medium_dp3')
self.add_free_floating_sbml(model_file=model_file, model_name='Medium_dp4')
cell_20 = self.fetch_cell_by_id(20)
self.add_sbml_to_cell(model_file=model_file, model_name='dp', cell=cell_20, integrator='gillespie')
def step(self, mcs):
self.timestep_sbml()
cell_20 = self.fetch_cell_by_id(20)
print('cell_20, dp=', cell_20.sbml.dp.values())
print('Free Floating Medium_dp2', self.sbml.Medium_dp2.values())
if mcs == 3:
Medium_dp2 = self.sbml.Medium_dp2
Medium_dp2['S1'] = 10
Medium_dp2['S2'] = 0.5
if mcs == 5:
self.delete_sbml_from_cell_ids(model_name='dp', cell_ids=list(range(1, 11)))
if mcs == 7:
cell_25 = self.fetch_cell_by_id(25)
self.copy_sbml_simulators(from_cell=cell_20, to_cell=cell_25)
In the start function we specify the path to the SBML model (here we use
partial path Simulation/test_1.xml because test_1.xml is in our CC3D
Simulation project directory) and also create a python dictionary that has
initial conditions for the SBML model. This particular model has two
floating species : S1 and S2 and our dictionary – initialConditions -
stores the initial concentration of these species to 0.0002 and 0.000002
respectively:
model_file = 'Simulation/test_1.xml'
initial_conditions = {}
initial_conditions['S1'] = 0.00020
initial_conditions['S2'] = 0.000002
Note
We can initialize each SBML Solver using different initial conditions. When we forget to specify initial conditions the SBML code usually has initial conditions defined and they will be used as starting values.
Before we discuss add_sbml_to_cell_ids function let us focus on statements
that open the start function:
options = {'relative': 1e-10, 'absolute': 1e-12}
self.set_sbml_global_options(options)
We set here SBML integrator options. These statements are optional,
however when your SBML model crashes with e.g. CVODE error, it often
means that your numerical tolerances (relative and absolute) or number
of integration steps in each integration interval (steps) should be
changed. Additionally you may want to enable stiff ODE solver by setting
stiff to True:
options = {'relative': 1e-10, 'absolute': 1e-12, 'stiff': False}
self.set_sbml_global_options(options)
After defining the options dictionary we inform CC3D to use these settings.
We do it by using as shown above. A thing to remember that new options
will apply to all SBML model that were added after calling
set_sbml_global_options. This means that usually you want to ensure that
SBML integration options setting should be the first thing you do in your
Python steppable file. If you want ot retrieve options simply type:
options = self.get_sbml_global_options()
Notice that options can be None indicating that options have not been set (this is fine) and the default SBML integrator options will be applied.
Let us see how we associate SBML model with several cells using add_sbml_to_cell_ids:
self.add_sbml_to_cell_ids(model_file=model_file, model_name='dp',
cell_ids=list(range(1, 11)), step_size=0.5, initial_conditions=initial_conditions)
This function looks relatively simple but it does quite a lot if you look under the hood. The first argument is the path to the SBML models file. The second one is the model alias - it is a name you choose for the model. It is an arbitrary model identifier that you use to access the model. The third argument is a Python list that contains cell ids to which CC3D will attach an instance of the SBML Solver.
Note
Each cell will get a separate SBML solver object. SBML Solver objects are associated with cells, while free floating SBML Solvers are independent.
The fourth argument specifies the size of the integration step – here we
use a value of 0.5 time unit. The fifth argument passes the optional initial conditions
dictionary.
Each SBML Solver function that associates models with a cell or adds a free floating model calls libRoadRunner functions that parse SBML and translate it to very fast LLVM code. Everything happens automatically and produces optimized solvers which are much faster than solvers that rely on some kind of interpreters.
The next five function calls to self.add_free_floating_sbml create instances of
SBML Solvers which are not associated with cells but, as you can see,
have distinct names. Unique naming of free floating models is required because
when we want to refer to such a solver to extract model values we will do so using the model name.
The reason all models attached to cells have same name is that when we
refer to such a model we pass the cell object and a name, which uniquely
identify the model. Notice that the last three calls to
self.add_free_floating_sbml do specify neither step size (we use the default step
size of 1.0 time unit) nor initial conditions (we use whatever defaults are
in the SBML code).
Finally, the last two lines of the start function demonstrates how to add an SBML Solver object to a single cell and select a non-default SBML Solver integrator:
cell_20 = self.fetch_cell_by_id(20)
self.add_sbml_to_cell(model_file=model_file, model_name='dp', cell=cell_20, integrator='gillespie')
Instead of a passing list of cell ids we pass a cell object (cell_20). All integrators
supported by libRoadRunner are available using the keyword integrator, which includes
CVODE (default), Gillespie (integrator='gillespie'), Euler (integrator='euler'),
Runge-Kutta (integrator='rk4') and Gillespie Direct Method (integrator='rk45').
For more information on the details of available integrators, visit the
libRoadRunner online documentation.
The integrator keyword argument is optional to, and available in, all functions
that create a SBML Solver instance.
We can also associate SBML model with certain cell types using the following syntax:
self.add_sbml_to_cell_types(model_file=model_file, model_name='dp', cell_types=[self.NONCONDENSING],
step_size=step_size, initial_conditions=initial_conditions)
This time instead of passing a list of cell ids we pass list of cell types.
Let us move on to the step function. The first call we see there is
self.timestep_sbml. This function carries out integration of all SBML
Solver instances defined in the simulation. The integration step can be
different for different SBML Solver instances (as shown in our example).
To check the values of model species after the integration step we can call e.g.
print('Free Floating Medium_dp2', self.sbml.Medium_dp2.values())
These functions check and print model variables for the free floating model
called Medium_dp2.
The next set of function calls:
if mcs == 3:
Medium_dp2 = self.sbml.Medium_dp2
Medium_dp2['S1'] = 10
Medium_dp2['S2'] = 0.5
set a new state for the free floating model called Medium_dp2. If we
wanted to print the state of the model dp belonging to the cell object called
cell_20 we would use the following syntax:
print('cell_20, dp=', cell_20.sbml.dp.values())
To assign new values to dp model variables for cell_20 we use the
following syntax:
cell_20.sbml.dp['S1'] = 10
cell_20.sbml.dp['S2'] = 0.5
Note
We access a free-floating SBML Solver via self.sbml.MODEL_ALIAS syntax whereas SBML Solvers associated with a particular cell are accessed using a reference to the cell objects e.g. cell_20.sbml.MODEL_ALIAS
Another useful operation within SBML Solver capabilities is deletion of models. This is handy when at a certain point in your simulation you no longer need to solve ODEs described in the SBML model. This is the syntax that deletes a named SBML Solver model from specific cell ids:
self.delete_sbml_from_cell_ids(model_name='dp', cell_ids=list(range(1, 11)))
As you probably suspect, we can delete a named SBML Solver model from cell types:
self.delete_sbml_from_cell_types(model_name='dp' ,cell_types=range[self.A,self.B])
from a single cell:
self.delete_sbml_from_cell(model_name='dp',cell=cell_20)
or delete a free floating SBML Solver object:
self.delete_free_floating_sbml(model_name='Medium_dp2'))
Note
When cells get deleted all SBML Solver models attached to them are deleted automatically. You do not need to call delete_sbml functions in such a case.
Sometimes you may encounter a need to clone all SBML models from one
cell to another (e.g. in the mitosis updateAttributes function where you
clone SBML Solver objects from a parent cell to a child cell). SBML Solver
lets you do that very easily:
cell_10 = self.fetch_cell_by_id(10)
cell_25 = self.fetch_cell_by_id(25)
self.copy_sbml_simulators(from_cell=cell_10, to_cell=cell_25)
What happens here is that source cell (from_cell) provides SBML Solver
object templates and based on these templates new SBML Solver objects
are gets created and CC3D assigns them to target cell (to_cell). All
the state variables in the target SBML Solver objects are the same as
values in the source objects.
If you want to copy only select models you would use the following syntax:
cell_10 = self.fetch_cell_by_id(10)
cell_25 = self.fetch_cell_by_id(25)
self.copy_sbml_simulators(from_cell=cell_10, to_cell=cell_25, sbml_names=['dp'])
As you can see there is a third argument - a Python list that specifies
which models to copy by name. Here we are copying only dp models. All other
models associated with the parent cells will not be copied.
This example demonstrates most important capabilities of SBML Solver.
The next example shows a slightly more complex simulation where we reset
initial condition of the SBML model before each integration step
(Demos/SBMLSolverExamples/DeltaNotch).
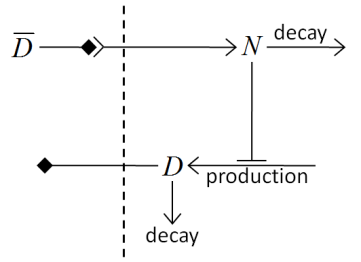
A full description of the Delta-Notch simulation is in the introduction to CompuCell3D Manual. The Delta-Notch example demonstrates multi-cellular implementation of Delta-Notch mutual inhibitory coupling. In this juxtacrine signaling process, a cell’s level of membrane-bound Delta depends on its intracellular level of activated Notch, which in turn depends on the average level of membrane-bound Delta of its neighbors. In such a situation, the Delta-Notch dynamics of the cells in a tissue sheet will depend on the rate of cell rearrangement and the fluctuations it induces. While the example does not explore the richness due to the coupling of sub-cellular networks with inter-cellular networks and cell behaviors, it already shows how different such behaviors can be from those of their non-spatial simplifications. We begin with the ODE Delta-Notch patterning model of Collier in which juxtacrine signaling controls the internal levels of the cells’ Delta and Notch proteins. The base model neglects the complexity of the interaction due to changing spatial relationships in a real tissue:
where \(D\) and \(N\) are the concentrations of activated Delta and Notch proteins inside a cell, respecively, \(\bar{D}\) is the average concentration of activated Delta protein at the surface of the cell’s neighbors, and \(a\) and \(b\) are saturation constants, and are Hill coefficients, and \(\nu\) is a constant that gives the relative lifetimes of Delta and Notch proteins.
Figure 18 Diagram of Delta-Notch feedback regulation between and within cells.
For the sake of simplicity let us assume that we downloaded the SBML model implementing the Delta-Notch ODEs. How do we use such SBML model in CC3D? Here is the code:
from random import uniform
from cc3d.core.PySteppables import *
class DeltaNotchClass(SteppableBasePy):
def __init__(self, frequency=1):
SteppableBasePy.__init__(self, frequency)
def start(self):
# adding options that setup SBML solver integrator
# these are optional but useful when encounteting integration instabilities
options = {'relative': 1e-10, 'absolute': 1e-12}
self.set_sbml_global_options(options)
model_file = 'Simulation/DN_Collier.sbml'
self.add_sbml_to_cell_types(model_file=model_file, model_name='DN', cell_types=[self.TYPEA], step_size=0.2)
for cell in self.cell_list:
dn_model = cell.sbml.DN
dn_model['D'] = uniform(0.9, 1.0)
dn_model['N'] = uniform(0.9, 1.0)
cell.dict['D'] = dn_model['D']
cell.dict['N'] = dn_model['N']
def step(self, mcs):
for cell in self.cell_list:
delta_tot = 0.0
nn = 0
for neighbor, commonSurfaceArea in self.get_cell_neighbor_data_list(cell):
if neighbor:
nn += 1
delta_tot += neighbor.sbml.DN['D']
if nn > 0:
D_avg = delta_tot / nn
cell.sbml.DN['Davg'] = D_avg
cell.dict['D'] = D_avg
cell.dict['N'] = cell.sbml.DN['N']
self.timestep_sbml()
In the start function we add SBML model (Simulation/DN_Collier.sbml) to
all cells of type A (it is the only cell type in this simulation besides
Medium). Later in the for loop we initialize D and N species from the
SBML model using random values so that each cell has a different SBML model starting
state. We also store the initial SBML model in a cell dictionary for
visualization purposes – see the full code in
Demos/SBMLSolverExamples/DeltaNotch. In the step function for each
cell we visit its neighbors and sum the value of Delta in the neighboring
cells. We divide this value by the number of neighbors (this gives the
average Delta concentration in the neighboring cells - D_avg). We pass
D_avg to the SBML Solver for each cell and then carry out integration for
the next time step. Before calling self.timestep_sbml we store the
values of Delta and Notch concentrations in the cell dictionary, but we
do it for visualization purposes only. As you can see from this
example SBML Solver programing interface is convenient to use, not to
mention SBML Solver itself is a very powerful tool which allows
coupling cell-level and sub-cellular scales.